Members involved:
Zoher GUEROUI, Emmanuelle MARIE, Christophe TRIBET,
Our ability to control biological processes at multiple scales is essential for understanding living systems in a quantitative manner. While many molecular players involved in cellular functions start to be discovered, numerous aspects of the processes controlling the functions of cells remain unknown. To examine these aspects, we are developing new methods aiming to control the spatiotemporal properties of either the environment of cells or biological processes at molecular and cellular scales:
– Magnetic control of biological processes
We are using magnetic technologies and nanoparticles to manipulate in space and time biomolecules and modified their regulation upon magnetic controls. As magnetic interactions are contactless, remote controlled, and deeply penetrate into thick materials, our approach can provide a novel strategy for controlling biological processes. For instance, we recently demonstrate the magnetic manipulation of mRNA conjugated to nanoparticles to spatially control translation-based protein synthesis.
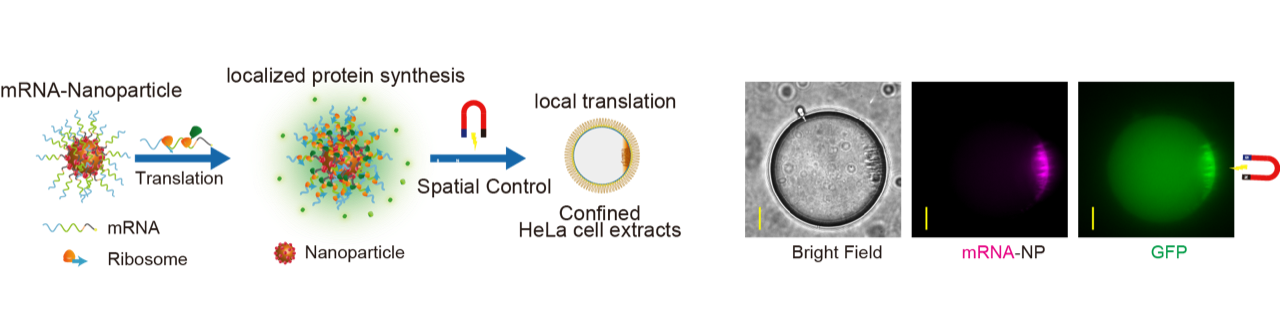
Left panel: Schematic illustrating messenger RNAs conjugated to iron-oxide nanoparticles that are designed as an RNA-based local source of protein synthesis. Right panel: Fluorescence microscopy observations of Cy5-labeled mRNA (Red) on nanoparticles colocalizing with translated GFP (Green).
Reference:
Nanoparticle-based local translation reveals mRNA as translation-coupled scaffold with anchoring function. Kashida S., Wang D.O., Saito H., and Gueroui Z. PNAS. 2019.
– Dynamic polymer coatings to control adhesion patterns
We implement spatiotemporal control of the presentation of chemical cues (ex adhesion peptides or proteins, repellant PEG, etc.) by stimuli-responsive layers of copolymers that are easy to use by non-specialists. Extending the poly(lysine)-based toolbox, we design light-, temperature-, or redox-triggered polymer switches, that spontaneously form layers on a variety of surfaces, and enable to modulate the interaction with cells or particles. We characterize the composition and responsiveness of mixed and/or micropatterned layers. We study soft-adhesion by micro-tracking of particles to model complex interactions. With living cells, we control time modulated adhesion/deadhesion (for instance on cadherin and integrin receptors). These layers open new possibilities for dynamic manipulation of cell polarization, or migration, and for cocultures.
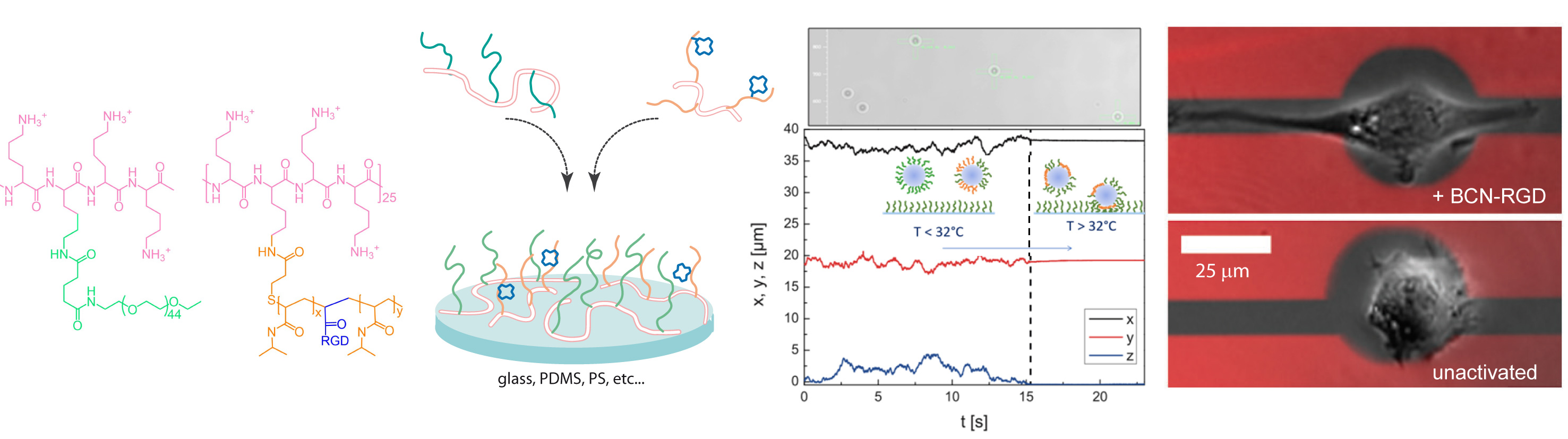
Tracking of colloid soft adhesion and control of cell binding & migration onto mixed layers of stimuli-responsive comb-like PLL derivatives.